LINKS ICaMB Mol Biol and Bioiformatics resources
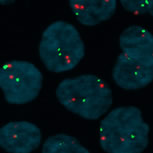
Interphase Fish showing the MLL locus on chromosome 11(red) and chromosome 11 centromeres (green) |
Ian Cowell
Institute for Cell and Molecular Biosciences Newcastle University E-mail i.g.cowell@ncl.ac.uk
Research Interests
Chromosomes, chromatin dynamics and genome stability. The eukaryotic cell nucleus is a highly organized but dynamic structure that is responsible for maintaining and faithfully replicating the genetic material. At the same time the nucleus is the site of transcription and RNA processing. How these processes are brought about and how they are coordinated are fundamental questions in modern biology with great importance to the understanding of conditions including cancer and ageing. Mechanisms of chromosomal translocation in leukaemia. Therapy-related leukaemias can occur in patients previously treated for a primary cancer and frequently display characteristic recurrent chromosome translocations. These translocations are thought to be causative or at least early contributory events for these leukaemias. How and why recurrent translocations occur is unknown, but the mechanism presumably involves chromosome breakage and incorrect rejoining. To better understand this process we are using interphase in situ hybridization (iFISH) and immunological techniques to probe the chromosome and chromatin dynamics of the regions involved. This work is carried out in Prof. Caroline Austin's laboratory as part of a Leukaemia Research Fund program grant. Changes in chromatin and nuclear organization associated with DNA damage and genome stability. The histone variant H2A.X is phosphorylated after cellular exposure to DNA-damaging agents such as ionizing radiation, resulting in chromatin domains enriched for phosphorylated H2A.X (gammaH2AX) around sites of double-strand DNA breaks. I previously showed that heterochromatin is a barrier to H2AX phosphorylation (10) and work subsequently published in Molecular Cell and Nature has confirmed this, showing that removal of the heterochromatin protein HP1 is a necessary part of the DNA damage response. Histone deacetylase inhibitors (HDACIs) cause histone hyperacetylation and alter the properties of heterochromatin relieving the block to H2AX phosphorylation. DNA topoisomerase II poisons such as etoposide are mainstream cytotoxic anticancer drugs, and recently it has been shown that their cytotoxic effects can be potentiated by HDACIs. In Professor Austin's laboratory we are interested in the interactions between histone deacetylase inhibitors, DNA topoisomerase II poisons and heterochromatin with the aim of better understanding the action of this class of drugs. Targeting DNA-repair proteins as a means to improve the efficacy of anti-cancer treatments. I contributed to work validating the PIKK kinases DNA-PKcs and ATM as drug targets for anti-cancer therapies (7,8 & 12). I am currently collaborating with Drs Durkacz and Wilmore in the NICR at Newcastle on an LRF-funded project to determine the role of DNA damage-inducible kinases in the cellular responses to nucleoside analogues used in leukaemia therapy. Histone lysine methylation and the function of chromodomain proteins. Work I started at Newcastle and then continued at the Babraham and the Roslin Institutes focused on histone modifications and the function of chromodomain proteins (2-4). Chromodomain proteins, particularly the heterochromatin protein HP1 bind to, and form the “readout” for modifications including trimethylation of histone H3 at lysine 9. I was amongst the first to report the evolutionarily widespread occurrence of chromatin domains enriched for histone H3 trimethylated at lysine 9, and the association of this epigenetic modification with heterochromatin and a transcriptionally silent state (5). Current work includes studying the relationship between the epigenetic status of genomic regions and their response to DNA damage induced by agents such as topoisomerase II poisons or ionizing radiation (see above).
|