ENZYME ASSEMBLIES & COMPLEXES
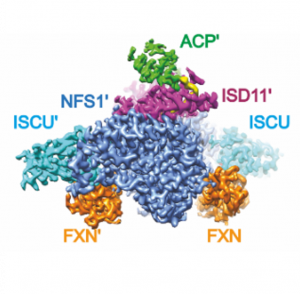
Many metabolic enzymes performing essential cellular roles have evolved to function as oligomeric assemblies, or in concert with other partner proteins. We have vested interest in studying metabolic enzyme complexes that are linked with genetic diseases. We adopt novel approaches in recombinant co-expression and endogenous isolation of metabolic complexes, for structural, biochemical and biophysical characterisation. Our latest work includes: frataxin-bound FeS assembly complex, glycogenin-glycogen synthase complex, and cystathionine beta-synthase oligomers.
INTEGRATIVE STRUCTURAL BIOLOGY
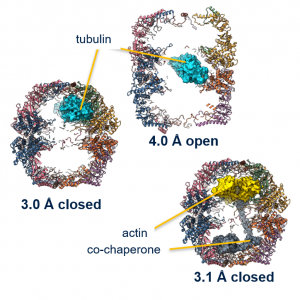
Our group combines x-ray crystallography, cryo-electron microscopy and complementary biophysical methods (including differential scanning fluorimetry, surface plasmon resonance, Michaelis-Mention kinetics) to study enzyme structures, dynamics and interactions. We have to date determined > 200 structures, all deposited in the Protein Data Bank. Our goal is to provide molecular insight into functional and disease-causing mechanisms. We recently applied cryo-EM to reveal conformation of client substrates and cochaperone within the TRiC chaperone.
SMALL MOLECULE DRUG DISCOVERY
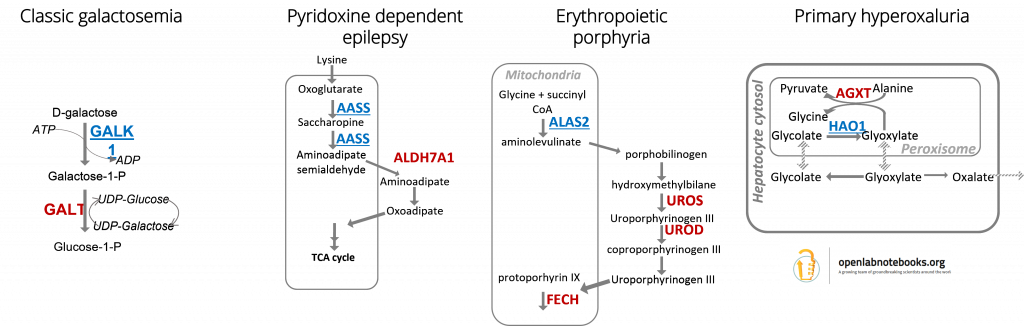
Disease-associated metabolic enzymes often lead to aberrant flux and accumulation of toxic metabolites. Therefore, pathway manipulation to reduce flux to the defective enzyme could have therapeutic benefit. Our objectives are to develop small molecule inhibitors for enzymes upstream of a metabolic defect, through the approach known as ‘substrate reduction therapy’. Check out our hit discovery projects of GALK1, HAO1, ALAS2 and AASS, in which our crystallography-based fragment screening campaigns are released in the public domain.